

I am a Research Associate and Lab Manager in a CAR-T cell Research Lab (email me for credentials specifics), and I find the ideas here very interesting. I will email GeneSmith to get more details on their research, and I am happy to provide whatever resources I can to explore this possibility.
TLDR;
Making edits once your editing system is delivered is (relatively) easy. Determining which edits to make is (relatively) easy. (Though you have done a great job with your research on this, I don't want to come across as discouraging.) Delivering gene editing mechanisms in-vivo, with any kind of scale or efficiency, is HARD.
I still think it may be possible, and I don't want to discourage anyone from exploring this further. I think the resources and time required to bring this to anything close to clinical application will be more than you are expecting. Probably on the order of 10-20 years, at least many millions (5-10 million?) of USD, in order to get enough data to prove the concept in mice. That may sound like a lot, but I am honestly not sure if I am being appropriately pessimistic. You may be able to advance that timescale with significantly more funding, but only to a poi...
Thanks for leaving such a high quality comment. I'm sorry for taking so long to get back to you.
We fully expect bringing this to market to take tens of millions of dollars. My best guess was $20-$40 million.
My biggest concern is your step 1:
"Determine if it is possible to perform a large number of edits in cell culture with reasonable editing efficiency and low rates of off-target edits."
And translating that into step 2:
"Run trials in mice. Try out different delivery vectors. See if you can get any of them to work on an actual animal."
I would like to hear more about your research into approaching this problem, but without more information, I am concerned you may be underestimating the difficulty of successfully delivering genetic material to any significant number of cells.
We expect this to be difficult, but we DON'T expect to have to solve the delivery problem entirely on our own. There are significant incentives for existing companies such as Dyno Therapeutics to solve the problem of delivering genes (or other payloads) to the nucleus of brain cells. In fact, Dyno already has a product, Dyno bCap 1 which successfully delivered genes to between 5% and 20% of brain cells in non-hum...
Really interesting, thanks for commenting.
My lab does research specifically on in vitro gene editing of T-cells, mostly via Lentivirus and electroporation, and I can tell you that this problem is HARD.
Even in-vitro, depending on the target cell type and the amount/ it is very difficult to get transduction efficiencies higher than 70%, and that is with the help of chemicals like Polybrene, which significantly increases viral uptake and is not an option for in-vivo editing.
Does this refer to the proportion of the remaining cells which had successful edits / integration of donor gene? Or the number that were transfected at all (in which case how is that measured)?
...Essentially, in order to make this work for in-vivo gene editing of an entire organ (particularly the brain), you need your transduction efficiency to be at least 2-3 orders of magnitude higher than the current technologies allow on their own just to make up for the lack of polybrene/retronecti
I'll give a quick TL;DR here since I know the post is long.
There's about 20,000 genes that affect intelligence. We can identify maybe 500 of them right now. With more data (which we could get from government biobanks or consumer genomics companies), we could identify far more.
If you could edit a significant number of iq-decreasing genetic variants to their iq-increasing counterpart, it would have a large impact on intelligence. We know this to be the case for embryos, but it is also probably the case (to a lesser extent) for adults.
So the idea is you inject trillions of these editing proteins into the bloodstream, encapsulated in a delivery capsule like a lipid nanoparticle or adeno-associated virus, they make their way into the brain, then the brain cells, and the make a large number of edits in each one.
This might sound impossible, but in fact we've done something a bit like this in mice already. In this paper, the authors used an adenovirus to deliver an editor to the brain. They were able to make the targeted edit in about 60% of the neurons in the mouse's brain.
There are two gene editing tools created in the last 7 years which are very good candidates for our task, with a low ...
Why would this work on adults? The brain develops most in childhood. If those genes' role is to alter the way synapses develop in the fastest growth phase, changing them when you're 30 won't do anything.
I mean, I explicitly state in the post that I don't think we'll be able to reach IQs far outside the normal human range by just flipping alleles:
I don’t expect such an IQ to actually result from flipping all IQ-decreasing alleles to their IQ-increasing variants for the same reason I don’t expect to reach the moon by climbing a very tall ladder; at some point, the simple linear model will break down.
So yes, I agree with you
Thinking about this post these days... Editing discussions might be better focused on personality: is that feasible, statistically? It seems like it might be, but we don't know.
The focus on IQ in older discussions strikes me as increasingly misguided. It's a good trait to start with, because it is important, well-studied, and turns out to be highly tractable, but it should only be a stepping stone to more useful approaches like index scores. There's also another reason to treat IQ as just a toy example: we are now well into the deep learning revolution, and it's come so far, and there's so much scope for scaling & improvement, that it seems like IQ is plummeting in value each year. Already it feels like people get less or more out of AI based on their flexibility and willingness to experiment or to step back & delegate & finish missing pieces. When the LLMs can do all the smart things you ask them to do, the value becomes in asking for good ones, and making good use of them. The future doesn't seem like it'll be kind to neurotic, eager-to-please types, but good to those who are unafraid to have clear visions or know what they want, finish projects and - pace Amdahl's la...
A key consideration when selecting for latent mental traits is whether a common pathway model holds for the latent variable under selection. In an ideal common pathway model, all covariance between indicators is mediated by a single underlying construct.
When this model fails, selecting for one trait can lead to unintended consequences. For instance, attempting to select for Openness might not reliably increase open-mindedness or creativity. Instead, such selection could inadvertently target specific parts of whatever went into the measurement, like liberal political values, aesthetic preferences, or being the kind of person with an inflated view of yourself.
Unlike personality factors, which demonstrate mixed evidence for a coherent latent structure, IQ has been more consistently modeled using a common pathway approach.
TL;DR: Selecting for IQ good. Will get smarter children. Selecting for personality risky. Might get child that likes filling in the rightmost bubble on tests.
Sources:
https://psycnet.apa.org/record/2013-24385-001
https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7839945/
I'm not sure we want something beyond statistical range of human personality traits
Obviously it is untrue that editing is useless if it 'only' gives you a von Neumann. Similarly for personality. We don't reify sets of personality traits as much as IQ, which is more obvious, but there are definitely many people who achieved remarkable things through force of personality. (Think figures like Lee Kuan Yew or Napoleon or Elon Musk come to mind as an example: they were smart, and lucky, and made good choices, but there is clearly still a lot left over to explain.) And because personality is many things and there seems to be a pipeline model of output, you quickly get very few people at the tails who assemble all the right components. (Gignac has a paper making this point more explicitly.)
Why do not select outliers from population using personality testing and give them high intelligence?
You're acting like it's uncontroversially true that you have unlimited edits and can change any property at any time in development. I don't think that is the case.* There is going to be an editing budget and limits to editing. One might as well ask the opposite question: why not select intelligen...
Using LLMs is an intellectual skill. I would be astonished if IQ was not pretty helpful for that.
I don't think it is all that helpful, adjusting for the tasks that people do, after years of watching people use LLMs. Smart people are often too arrogant and proud, and know too much. "It's just a pile of matrix multiplications and a very complicated if function and therefore can't do anything" is the sort of thing only a smart person can convince themselves, where a dumb person thinking "I ask the little man in the magic box my questions and I get the right answers" is getting more out of it. (The benefits of LLM usage is also highly context dependent: so you'll find studies showing LLMs assist most the highest performers, but also ones showing it helps most the lowest.) Like in 2020, the more you knew about AI, the dumber your uses of GPT-3 were, because you 'knew' that it couldn't do anything and you had to hold its hand to do everything and you had to phrase everything in baby talk etc. You had to unlearn everything you knew and anthropomorphize it to meaningfully explore prompting. This requires a certain flexibility of mind that has less to do with IQ and more to do with, say, schizophrenia - the people in Cyborgism, who do the most interesting things with LLMs, are not extraordinarily intelligent. They are, however, kinda weird and crazy.
I remain both skeptical some core claims in this post, and convinced of its importance. GeneSmith is one of few people with such a big-picture, fresh, wildly ambitious angle on beneficial biotechnology, and I'd love to see more of this genre.
One one hand on the object level, I basically don't buy the argument that in-vivo editing could lead to substantial cognitive enhancement in adults. Brain development is incredibly important for adult cognition, and in the maybe 1%--20% residual you're going well off-distribution for any predictors trained on unedited individuals. I too would prefer bets that pay off before my median AI timelines, but biology does not always allow us to have nice things.
On the other, gene therapy does indeed work in adults for some (much simpler) issues, and there might be valuable interventions which are narrower but still valuable. Plus, of course, there's the nineteen-ish year pathway to adults, building on current practice. There's no shortage of practical difficulties, but the strong or general objections I've seen seem ill-founded, and that makes me more optimistic about eventual feasibility of something drawing on this tech tree.
I've been paying clo...
ANNs and BNNs operate on the same core principles; the scaling laws apply to both and IQ in either is a mostly function of net effective training compute and data quality. Genes determine a brain's architectural prior just as a small amount of python code determines an ANN's architectural prior, but the capabilities come only from scaling with compute and data (quantity and quality).
So you absolutely can not take datasets of gene-IQ correlations and assume those correlations would somehow transfer to gene interventions on adults (post training in DL lingo). The genetic contribution to IQ is almost all developmental/training factors (architectural prior, learning algorithm hyper params, value/attention function tweaks, etc) which snowball during training. Unfortunately developmental windows close and learning rates slow down as the brain literally carves/prunes out its structure, so to the extent this could work at all, it is mostly limited to interventions on children and younger adults who still have significant learning rate reserves.
But it ultimately doesn't matter, because the brain just learns too slowly. We are now soon past the point at which human learning matters much.
Regulation and complexity of effects seem like another two big blockers.
Effects of genes are complex. Knowing a gene is involved in intelligence doesn't tell us what it does and what other effects it has.
I wouldn't accept any edits to my genome without the consequences being very well understood (or in a last-ditch effort to save my life). I'd predict severe mental illness would happen alongside substantial intelligence gains.
Source: research career as a computational cognitive neuroscientist.
I put this as a post- ASI technology, but that's also a product of my relatively short timelines.
Yes, I think many in the field would share this viewpoint and that's part of why we haven't seen someone already attempt this.
I disagree for reasons I've shared in my post on "Black Box Biology", but it's worth reiterating my reasons here:
Your point 2 is my big hangup. If you mean each genetic variant affects each trait roughly linearly, sure. If you mean each genetic variant affects only one trait, I think that's completely wrong. Most studies only address the affect on a single trait, but given the re-use of proteins for different roles, I fully expect multiple effects of genes on average. It seems like I've seen studies and papers on this, but it's never been my area, so I don't remember anything clearly.
Evolution succeeds by tinkering over many generations. It creates as many downsides as upsides. Who's going to volunteer to be tinkered upon?
Actually, as soon as I pose the question that way, I realize that the answer is "lots of people" (as long as there's a reason to think you'll get more upside than downside, which limited theory will provide.)
However, there's no way the FDA is going to approve tinkering. You'd have to do this outside of US jurisdiction.
I am not saying plieotropy doesn't exist. I'm saying it's not as big of a deal as most people in the field assume it is.
Take disease risk for example. Here's a chart showing the genetic correlations between various conditions:
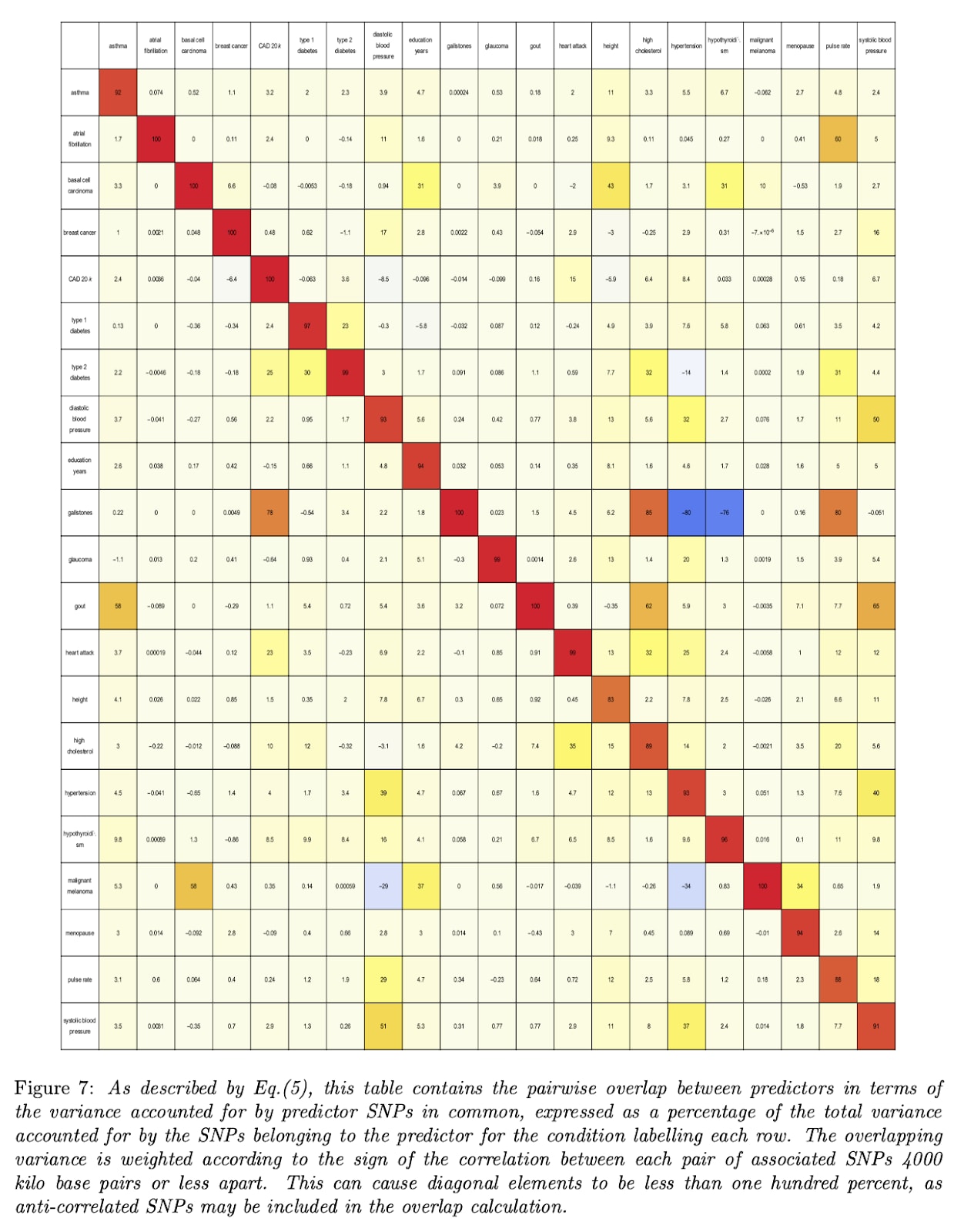

With a few notable exceptions, there is not very much correlation between different diseases.
And to the extent that plieotropy does exist, it mostly works in your favor. That's why most of the boxes are yellowish instead of bluish. Editing or selecting embryos to reduce the risk of one disease usually results in a tiny reduction of others.
Evolution succeeds by tinkering over many generations. It creates as many downsides as upsides. Who's going to volunteer to be tinkered upon?
Evolution cannot simultaneously consider data from millions of people when deciding which genetic variants to give someone. We can.
None of these proposals deal with novel genetic variants. Every target variant we would introduce is already present in tens of thousands of individuals and is known to not cause any monogenic disorder.
as long as there's a reason to think you'll get more upside than downside, which limited theory will provide
I'm not quite sure what you're getting at here. Do ...
Me: "I don't think this therapy as OP describes it is possible for reasons that have already been stated by HiddenPrior and other reasons"
kman: "Can you elaborate on this? We'd really appreciate the feedback."
Considering the enormity of my response, I figured I would post it in a place that is more visible to those interested. First I'd like to express my gratitude for you and GeneSmith's goal and motivation; I agree that without some brain-machine interface solution, intelligence enhancement is certainly the way forward for us if we'd like to not only keep up with AI, but also break through the intellectual soft caps that have led to plateauing progress in many fields as they gradually become hyper-specialized. I don't use this website very often, but fortunately this post was sent to me by a friend and I decided to engage with it since I had the exact same ideas and motivations detailed in your post when I matriculated into a PhD program for molecular genetics. I don't want to de-anonymize myself, so I won't mention the exact school but it is ranked within the top 50 graduate programs in the US and has had many Nobel Prize laureates as faculty. I was very optimistic, like you two...
You could elect to use proxy measures like educational attainment, SAT/ACT/GRE score, most advanced math class completed, etc., but my intuition is that they are influenced by too many things other than pure g to be useful for the desired purpose. It's possible that I'm being too cynical about this obstacle and I would be delighted if someone could give me good reasons why I'm wrong.
This is just measurement error and can be handled by normal psychometric approaches like SEM (eg. GSEM). You lose sample efficiency, but there's no reason you can't measure and correct for the measurement error. What the error does is render the estimates of each allele too small (closer to zero from either direction), but if you know how much error there is, you can just multiply back up to recover the real effect you would see if you had been able to use measurement with no error. In particular, for an editing approach, you don't need to know the estimate at all - you only need to know that it is non-zero, because you are identifying the desired allele.
So, every measurement on every individual you get, whether it's EA or SAT or GRE or parental degree or a 5-minute web quiz, helps you narrow down th...
The problem could potentially be solved by conducting GWASes that identify the SNPs of things known to correlate with the proxy measure other than intelligence and then subtracting those SNPs
More or less. If you have an impure measurement like 'years of education' which lumps in half intelligence and half other stuff (and you know this, even if you never have measurements of IQ and EDU and the other-stuff within individuals, because you can get precise genetic correlations from much smaller sample sizes where you compare PGSes & alternative methods like GCTA or cross-twin correlations), then you can correct the respective estimates of both intelligence and other-stuff, and you can pool with other GWASes on other traits/cohorts to estimate all of these simultaneously. This gets you estimates of each latent trait effect size per allele, and you just rank and select.
you will most likely have the majority of causal intelligence SNPs falling below the genome-wide significance threshold of p < 5 * 10E-8.
A statistical-significance threshold is irrelevant NHST mumbo-jumbo. What you care about is posterior probability of the causal variant's effect being above the cost-safety threshold, whatever that may be, but which will have nothing at all to do with 'genome-wide statistical significance'.
Genetically altering IQ is more or less about flipping a sufficient number of IQ-decreasing variants to their IQ-increasing counterparts. This sounds overly simplified, but it’s surprisingly accurate; most of the variance in the genome is linear in nature, by which I mean the effect of a gene doesn’t usually depend on which other genes are present.
So modeling a continuous trait like intelligence is actually extremely straightforward: you simply add the effects of the IQ-increasing alleles to to those of the IQ-decreasing alleles and then normalize the score relative to some reference group.
If the mechanism of most of these genes is that their variants push something analogous to a hyperparameter in one direction or the other, and the number of parameters is much smaller than the number of genes, then this strategy will greatly underperform the simulated prediction. This is because the cumulative effect of flipping all these genes will be to move hyperparameters towards optimal but then drastically overshoot the optimum.
I'd be worried about changes to my personality or values from editing so many brain relevant genes.
Consolidating my previous comments:
I discussed this project with GeneSmith and I think it is promising, though very challenging to implement in practice. The hardest part will be safely and efficiently delivering the editing agent to a large fraction of the cells in the brain.
Some other points:
CAR T-cell therapy, a treatment for certain types of cancer, requires the removal of white blood cells via IV, genetic modification of those cells outside the body, culturing of the modified cells, chemotherapy to kill off most of the remaining unmodified cells in the body, and reinjection of the genetically engineered ones. The price is $500,000 to $1,000,000.
And it only modifies a single gene.
This makes it sound like CAR-T is gene editing, but it isn't. Instead of editing a gene, it introduces a new one (a chimeric T-cell receptor). Although some companies are working on gene editing to enhance CAR-Ts.
I also know of a PHD student in George Church’s lab that was able to make several thousand edits in the same cell at the same time by targeting a gene that has several thousand copies spread throughout the genome.
The paper reporting this was here: https://www.ncbi.nlm.nih.gov/pmc/article...
Thanks! I've added a link to the paper by Cory Smith and others to the relevant section and updated the section about CAR T-cell therapy to read "adds a single gene" rather than "modifies a single gene"
Can you provide an update on the current status and future plans for this idea? Where has it progressed so far, what direction is it expected to take moving forward?
I've started a gene therapy company, raised money, opened a lab, hired the inventor of one of the best multiplex gene editing techniques to be our chief scientific officer, and am currently working on cell culture experiments with the help of a small team.
I may write a post about what's happened at some point. But things are moving.
None of the stuff that you suggested has worked for any animal. I'm not saying it's impossible, but it is far harder to achiever compared to the stuff that HAS been demonstrated on mice.
I am PhD in Bio, have an extensive experience with stem cells and gene editing. The idea of human/animal cognitive enhancement is great, but the delivery of gene therapy to adult brains is doomed: first, it's technically challenging if not impossible, second, if we want to achieve a true revolution in cognition, we need to target brain development not already developed brain!
Imagine a monkey thinking of enhancing its abilities by injecting virus in its brain - will it ever reach a human level cognition? Sounds laughable. Who cares about +5 points to IQ, I want to see a 10xEinstein ;)
For >40 years, way before the discovery of CRISPRs and base editors, we've been successfully genetically engineering mice, but not other species. Why only mice? Because we can culture mouse embryonic stem cells that can give rise to complete animals. We did not understand why mouse cells were so developmentally potent, and why this didn't work for other species. Now we do (I'm the last author):
Highly cooperative chime...
Using tetraploid complementation, it is possible to achieve up to 70% of full-term development, which is similar rate of mouse natural conception. And this was before we understood how it works. I believe that soon we will be able to outperform nature and achieve close to 100% full term development and survival (I've seen 90% efficiency in some experiments). For human, only 30% of naturally conceived embryos are born, and only 10% of IVF, so superseding nature for human will be even easier than for mice.
In figure 4 we demonstrate that the mice are healthy, and can breed giving rise to healthy progeny, which is the highest bar for the quality of the cells. Again, our current IVF practice has only 10% s...
Thanks, this is very interesting.
One thing I don't understand: you write that a major problem with viruses is:
As one might expect, the immune system is not a big fan of viruses. So when you deliver DNA for a gene editor with an AAV, the viral proteins often trigger an adaptive immune response. This means that when you next try to deliver a payload with the same AAV, antibodies created during the first dose will bind to and destroy most of them.
Is this a problem for people who expect to only want one genetic modification during their lifetime?
So there are two separate concerns:
One is a concern for people who are getting a single dose monogenic gene therapy who already have antibodies to an AAV delivery vector due to a natural infection. In these cases, doctors can sometimes switch the therapy to use an AAV with a different serotype that can't be attacked by the patient's existing antibodies. If that's not available, they'll sometimes give patients immunosupressants.
The problem is more relevant in the context of multiplex editing because you may not be able to make all the edits you'd like to in one round of therapy. You can only inject so many AAVs or lipid nanoparticles or what have you at a time. Cells only have a limited capacity to process and break down editor proteins, and other waste products of the editing process. So you may need to do multiple editing rounds to achieve the desired effect.
It will be a lot easier to do this if the delivery vector itself doesn't trigger the immune system. If it does, antibodies formed during the first round of edits will attack and destroy the the delivery vector. Maybe you can just give someone immunosupressants during each round of treatment? Or maybe you can just use a different AAV? There are potential solutions but I don't yet know which are likely to work best.
If anyone is interested, I'm working on conducting small-scale, informal random and controlled trials for various nootropics that improve neuroplasticity and IQ, from dihexa to anthocyanin. The medicine I want to test has documented effects, but I want to gather more data on what mental subskills they improve performance on the most and how strong the effect is.
In order to aid recruitment, every participant will get to take the intervention, just in a staggered order.
I have several people who've agreed to participate. However, I've paused the initiative to focus on other projects. Let me know if you want to take over as lead researcher! The mesh will thank you.
```
group 1 group 2 group 3
phase 1:active control control
phase 2:control control active
phase 3:control active active
```If you get massively multiplex editing working in a dish, there are a bunch of studies these days exposing neurons in a dish to stimuli and training them to play some game or do some task.
Imagine if over 100 dishes of edited neurons you saw a slight (statistically significant) increase in performance.
Also, don't sleep on the astrocytes.
It is the great paradox of this forum to serve both as a platform where individuals aligned with the effective altruism movement raise alarms and invite deep reflection on existential risks—usually with a nuanced, subtle, and cautious approach to these topics—and as a space where individuals adhering to libertarian or transhumanist ideologies also promote rather radical ideas, which themselves might constitute new existential challenges for humanity. I say this not with the intent of disparagement but rather as an observation.
This topic is a fascinating example of this paradoxical mixture. On one hand, the author seems to appeal to fears surrounding the "superintelligence" of artificial intelligence to, in a way, justify the development of a "superintelligence" in humans, or at least a significant enhancement of intelligence for a select group of individuals—an elite of scientists somewhat reminiscent of the "Manthan project," aimed at solving the alignment problem. These would be heroes of humanity, capable of mastering the hydra of AI in a titanic intellectual struggle.
In reality, I think everyone here is aware that this argument is primarily rhetorical, as the risk associated wi...
You are right. When I wrote my initial comment, I believed the argument was self-evident and did not require elaboration. However, "self-evidence" is not an objective concept, and I likely do not share the same socio-cultural environment as most users of this platform. Upon reading your comment and Ben Pace's, I realize that this apparent self-evidence is far from universally shared and requires further explanation. I have already expanded on my argument in my previous response, but here are the specific reasons why I think the author's project (and indeed the transhumanist project of enhancing human beings) raises unprecedented issues in terms of increasing inequality, more so than most technological innovations such as running water or mobile phones.
First, John Rawls's veil of ignorance constitutes a strong philosophical and rational argument for considering excessive inequalities as unjust and morally condemnable (edit : this is not my personal claim but that of John Rawls, with whom I fully agree). This veil of ignorance aligns utilitarianism with Kant morality, as it invites the moral agent to step outside their specific case and evaluate the morality of a situation from a mor...
Could someone open a manifold market on the relevant questions here so I could get a better sense of the probabilities involved? Unfortunately, I don't know the relevant questions or the have the requisite mana.
Personal note- the first time I came into contact with adult gene editing was the youtuber Thought Emporium curing his lactose intolerance, and I was always massively impressed with that and very disappointed the treatment didn't reach market.
Why hasn't someone already done this?
I think this is a very important question to ask.
Sometimes the reasons are encouraging:
Other times the reasons are discouraging:
These definitely aren't exhaustive lists. It's just what came to me after a few minutes of thinking.
I see this as a sort of "stand on the shoulders of giants" type of situation. By consulting with domain experts and figuring out why people currently aren't doing this, you're standing on their shoulders. Once you're up there and have a better view of the path forward:
I'm glad to see that you've put some solid thought into this question. However, I was concerned to see this:
...But I’ve only talked to a coup
Promoted to curated: I am quite excited about this. If indeed adult gene editing is possible, this does open up a huge range of possibilities, and seems like a very important development to track. This post is also quite well written, and explains a bunch of important concepts around gene editing and intelligence and the relationship between genes and high-level traits of an organism pretty well.
There's a great youtuber called the thought emporium who did genetic engineering on himself. I highly recommend checking them out :
https://www.youtube.com/watch?v=J3FcbFqSoQY
And the 2year follow up: https://www.youtube.com/watch?v=aoczYXJeMY4
The tldr is he created a virus then ate it to make his digestive system have more of the gene that makes lactase as he was very intolerant. 2 years later the effects are starting to wear off as cells get replaced but it seems to have had a very high ROI
If you unintentionally insert a few base pairs into the promoter region of some protein, will the promoter just work a little less well, or will it break altogether? I
Molecular Biologist here. Promoters (and any non-coding regulatory sequence for that matter) are extremely sensitive to point mutations. Since their sequence determines how well the RNA-polymerase binds to them, any change in the sequence of bind motifs or even in the distance between these motifs has a major (generally negative) impact in transcription initiation efficiency. https://www.natu...
I am a relatively recently reformed geneticist/molecular biologist and previously used CRISPR/Cas9 at the bench in an experimental context. I no longer work in the lab and admit am not well-read on the latest literature.
I think this approach is interesting, and theoretically executable, but practically infeasible at the current maturity level of the relevant technologies. I’m not sure such a mission would be a good use of expertise and money at this stage. I share the views of a lot of the top level commenters here about the limited feasibility of the appr...
Good post. This looks possible, if not feasible.
"crazy, unpredictable, and dangerous" are all "potentially surmountable issues". It's just that we need more research into them before they stop being crazy, unpredictable, and dangerous. (except quantum I guess)
I think that most are focusing on single-gene treatments because that's the first step. If you can make a human-safe, demonstrably effective gene-editing vector for the brain, then jumping to multiplex is a much smaller step (effective as in does the edits properly, not necessarily curing a disease). ...
A huge and fascinating topic. But... I find myself thinking: suppose I wanted to change the color of my eyes. I could figure out how to gene-hack my iris - or I could get colored contact lenses.
If the objective is to make people smarter, to what extent can this be accomplished by being specific about the cognitive skills that are to be enhanced, and then identifying an appropriate set of tools?
This is obviously not a very realistic model, but it probably produces fairly realistic results. But again, this is an area for future improvement.
Curious from a modeling perspective: what improvements would be top of mind for you? Another way to phrase this: if someone else were to try modeling this, what aspects would you look at to tell if it's an improvement or not?
Hey, I found this super interesting and looks like you've done your research. I know the good folks over at IndieBio and I think you are at a really good stage to have a chat with them. I think they could be helpful in helping you find a co-founder and initial and follow-on funding for the company. I'm sure there are a number of other synthetic bio accelerators that would be helpful as well, but I have personal experience with them.
This is an interesting post, but it has a very funny framing. Instead of working on enhancing adult intelligence, why don't you start with:
Editing the brains of adult humans and expecting intelligence enhancement is like 3-4 impossibiliti...
Some points:
I read of a proposal a few months back to achieve brain immortality via introduction of new brain tissue that can be done in a way as to maintain continuity of experience and personality over time. Replenisens , Discussion on a system for doing it in human brains That would perhaps provide a more reliable vector for introduction, as the brain is progressively hybridised with more optimal neural genetic design. Perhaps this could be done more subtly via introduction of 'perfected' stem cells and then some way of increasing rate of die off of ol...
Evidence that adult cognition can be improved is heartening. I'd always had a small amount of fear regarding being "locked in" to my current level of intelligence with no meaningful scope for improvement. Long ago, in a more naive age, it was the prospect of children being enhanced to leave their parents in the dirt. Now, it looks like AI is improving faster than our biotechnology is.
It's always a pleasure to read deep dives into genetic engineering, and this one was uniquely informative, though that's to be expected from GeneSmith.
It seems plausible to me that some portion of iq-enhancing genes work through pathways outside the brain (blood flow, faster metabolism, nutrient delivery, stimulant-like effects, etc.). If that is the case and even just a small portion of the edits don't need to make it to the brain, couldn't you get huge iq increases without ever crossing the blood-brain barrier?
If timelines are short, it seems worthwhile to do that first and use the gains to bootstrap from there. Would that give significant returns fast enough to be worth doing? Is this something you're already trying to do?
Since I am not good enough to use English as my native language, I am writing this with the help of a translator. Is there a specific procedure that I should follow to participate as a subject in this treatment? I also have a low intelligence, so many parts of my life are painful, so if I could pursue a better life by receiving these technological benefits and doing what I want, it would be great. Although I do not have a decent degree, I hope that I can be of help to your research as much as possible as a subject.
I'm sure you've already thought of this, and I know nothing about this area of biology, but isn't it possible that the genes coding for intelligence more accurately code for the developmental trajectory of the brain, and that changing them would not in fact affect an adult brain?
Future readers of this post might be interested this other lesswrong post about the current state of multiplex gene editing: https://www.lesswrong.com/posts/oSy5vHvwSfnjmC7Tf/multiplex-gene-editing-where-are-we-now
The LessWrong Review runs every year to select the posts that have most stood the test of time. This post is not yet eligible for review, but will be at the end of 2024. The top fifty or so posts are featured prominently on the site throughout the year.
Hopefully, the review is better than karma at judging enduring value. If we have accurate prediction markets on the review results, maybe we can have better incentives on LessWrong today. Will this post make the top fifty?
Thanks for posting! From a ten thousand foot non-expert perspective I always sort of assumed there wouldn't be much extra challenge to editing multiple genes at once relative to editing a single gene - if you're deploying something to go in and make edits to every cell on the body, why can't it make more than one edit? Or failing that why can't multiple editing agents be in the same treatment? Interesting to read about the practical challenges and solutions.
Naive question : about immunogenicity, what are the problems with the obvious strategy to counter it ? (target the thymus first to "whitelist" the delivery method).
This is a very fun post, and I wish you all the best. To be briefly negative, it seems more or less completely impossible for this to impact AI outcomes with any reasonable model of the world at all, because it will not provide enough enhancement quickly enough. But it's a cool direction, and worth pursuing as its own science!
This seems great. Better to have superintelligent people than superintelligent AI.
I only have one concern, which is how the IQ is measured? I was assessed as a child and told I was average IQ and that I'd probably never learn English or Mathematics. But when I was 12 I scored 125 on the Mensa.dk test, and a while ago I scored 150+ on BRGHT so I'm not as dumb as the system believes. Not that I would recommend making any more of me.
That type of IQ tests only measure one type of intelligence, and I believe that verbal intelligence is more important for regula...
What's your best guess for what percentage of cells (in the brain) receive edits?
Are edits somehow targeted at brain cells in particular or do they run throughout the body?
CAR T-cell therapy, a treatment for certain types of cancer, requires the removal of white blood cells via IV, genetic modification of those cells outside the body, culturing of the modified cells, chemotherapy to kill off most of the remaining unmodified cells in the body, and reinjection of the genetically engineered ones. The price is $500,000 to $1,000,000.
And it only modifies a single gene.
This makes it sound like CAR-T is gene editing, but it isn't. Instead of editing a gene, it introduces a new one (a chimeric T-cell receptor). Although some c...
OP, are the rumors true? Will you still be working on enhancing adult intelligence as opposed to editing embryos?
Quite confused about the non-coding region edit hypothesis.
Either you mean "non-coding" as in "regulatory" in which case... wouldn't off-target mutation be just as bad?
Or do you mean "non-coding" as in "areas with an undetermined role that we currently assume are likely vestigial" - in which case, wouldn't the therapy have no effect since the regions aren't causal to anything, just correlated? [Or, in the case where I'd have an effect, we ought to assume that those "non-coding" regions are quite causal for many things and thus just as dangerous to edit]
great article. I hope you realize your startup research/idea. One comment, I think the salaries derail the whole budget plan, afaik from startup world I have been involved, founders make big sacrifices to get their thing going in return for a big equity in the startup they believe someday will become a unicorn.
This is super interesting and I have a question:
How difficult would it be to also apply this to the gamates and thus make any potential offspring also have the same enhanced intelligence (but this time it would go into the gene pool instead of just staying in the brain)? Does the scientific establishment think this is ethical? (Also, if you do something like this, you reduce the homogeneity of the gene pool which could make the modified babies very susceptible to some sort of disease. Would it be worth it to give the GMO babies a random subset of the changes to increase variation?)
Will you be working on enhancement of other cognitive abilities besides intelligence, such as memory?
We know that some genes are only active in the womb, or in childhood, which should make us very skeptical that editing them would have an effect.
Would these edits result in demethylated DNA? A reversion of the epigenome could allow expression of infant genes. There may also be robust epigenomic therapies developed by the time this project would be scalable.
Companies like 23&Me genotyped their 12 millionth customer two years ago and could probably get at perhaps 3 million customers to take an IQ test or submit SAT scores.
Just as you mentioned acade...
I see the biggest problem not on the technical side of things, but on the social side. The existing power balance withing the population and the fact that it discourages cooperation is in my opinion a much bigger obstacle to alignment. Heck, it prevents alignment between human groups, let alone between humans and the future AGI. I don't see how increased intelligence of a small select group of humans can solve this problem. Well, maybe I am just not smart enough.
These are merely my qualifications I would place on this seemingly very interesting and highly well informed, well researched and competent idea the author has raised. Overall, these qualifications are of the "meta" sociological and political intersections onto the trait being enhanced here. Before presenting them, I would also like to state that the novel characteristics and approach to this subject that the author, who is not formally educated in biology and free of its academic milieu, is suggestive that the culture that surrou...
I took the pfizer vaccine and I found the section explaining how the mRNA vaccine works both illuminating and disturbing. Getting cells to generate and display the spike element of the virus, causing inflammation - and potentially cell death is perhaps all fine in the muscle tissue of the shoulder, but what happens if the mRNA travels further to different tissue like the heart? Upon reading how the vaccine actually works I have become a lot more suspicious of potential bad effects of the vaccine - previously I was under the naive impression that the vaccin...
Man, if you think the vaccine is scary just wait until you hear about what COVID does.
In all seriousness, you shouldn't find mRNA vaccines any scarier than the J&J vaccine or the Novavax vaccine. J&J puts DNA for the spike protein in a modified adenovirus, which then enters your muscle cells, breaks out into the cytoplasm, and injects the DNA payload into the nucleus. Then RNA transcriptase makes mRNA out of that DNA, which is exported from the nucleus into the cytoplasm where it is turned into the spike protein.
Novavax just skips the entire manufacturing process; they just inject the spike protein directly into your body.
I'm not an expert on COVID vaccines, but from everything I have read, the rare dangerous side-effects like myocarditis seem to come from the spike protein itself triggering a dangerous immune response in some very small percentage of people.
But like... you can't avoid that with any vaccine. And the incident of getting myocarditis or a worse condition from COVID itself (especially if you're unvaccinated) are way, way higher than getting it from the vaccine.
Vaccines are mostly just a cost-benefit analysis; if your odds of damage are higher from the virus (ad...
TL;DR version
In the course of my life, there have been a handful of times I discovered an idea that changed the way I thought about where our species is headed. The first occurred when I picked up Nick Bostrom’s book “superintelligence” and realized that AI would utterly transform the world. The second was when I learned about embryo selection and how it could change future generations. And the third happened a few months ago when I read a message from a friend of mine on Discord about editing the genome of a living person.
We’ve had gene therapy to treat cancer and single gene disorders for decades. But the process involved in making such changes to the cells of a living person is excruciating and extremely expensive. CAR T-cell therapy, a treatment for certain types of cancer, requires the removal of white blood cells via IV, genetic modification of those cells outside the body, culturing of the modified cells, chemotherapy to kill off most of the remaining unmodified cells in the body, and reinjection of the genetically engineered ones. The price is $500,000 to $1,000,000.
And it only adds a single gene.
This is a big problem if you care about anything besides monogenic diseases. Most traits, like personality, diabetes risk, and intelligence are controlled by hundreds to tens of thousands of genes, each of which exerts a tiny influence. If you want to significantly affect someone’s risk of developing lung cancer, you probably need to make hundreds of edits.
If changing one gene to treat cancer requires the involved process described above, what kind of tortuous process would be required to modify hundreds?
It seemed impossible. So after first thinking of the idea well over a year ago, I shelved it without much consideration. My attention turned to what seemed like more practical ideas such as embryo selection.
Months later, when I read a message from my friend Kman on discord talking about gene editing adults in adult brains, I was fairly skeptical it was worth my time.
How could we get editing agents into all 200 billion brain cells? Wouldn’t it cause major issues if some cells received edits and others didn’t? What if the gene editing tool targeted the wrong gene? What if it provoked some kind of immune response?
But recent progress in AI had made me think we might not have much time left before AGI, so given that adult gene editing might have an impact on a much shorter time scale than embryo selection, I decided it was at least worth a look.
So I started reading. Kman and I pored over papers on base editors and prime editors and in-vivo delivery of CRISPR proteins via adeno-associated viruses trying to figure out whether this pipe dream was something more. And after a couple of months of work, I have become convinced that there are no known fundamental barriers that would prevent us from doing this.
There are still unknowns which need to be worked on. But many pieces of a viable editing protocol have already been demonstrated independently in one research paper or another.
If this works, it’s going to be a very big deal
It is hard to overstate just how big a deal it would be if we could edit thousands of genes in cells all over the human body. Nearly every trait and every disease you’ve ever heard of has a significant genetic component, from intelligence to breast cancer. And the amount of variance already present in the human gene pool is stunning. For many traits, we could take someone from the 3rd percentile to the 97th percentile by editing just 20% of the genes involved in determining that trait.
Tweaking even a dozen genes might be able to halt the progression of Alzheimer’s, or alleviate the symptoms of major autoimmune diseases.
The same could apply to other major causes of aging: diabetes, heart disease, cancers. All have genetic roots to some degree or other. And all of this could potentially be done in people who have already been born.
Of particular interest to me is whether we could modify intelligence. In the past ten thousand years, humans have progressed mostly through making changes to their tools and environment. But in that time, human genes and human brains have changed little. With limitations to brain power, new impactful ideas naturally become harder to find.
This is particularly concerning because we have a number of extremely important technical challenges facing us today and a very limited number of people with the ability to make progress on them. Some may simply be beyond the capabilities of current humans.
Of these, the one I think about most is technical AI alignment work. It is not an exaggeration to say that the lives of literally everyone depend on whether a few hundred engineers and mathematicians can figure out how to control the machines built by the mad scientists in the office next door.
Demis Hassabis has a famous phrase he uses to describe his master plan for Deepmind: solve intelligence, then use that intelligence to solve everything else. I have a slightly more meta level plan to make sure Demis’s plan doesn’t kill everyone; make alignment researchers smarter, then kindly ask those researchers to solve alignment.
This is quite a grandiose plan for someone writing a LessWrong blog post. And there’s a decent chance it won’t work out for either technical reasons or because I can't find the resources and talent to help me solve all the technical challenges. At the end of the day, whether or not anything I say has an impact will depend on whether I or someone else can develop a working protocol.
How does intelligence even work at a genetic level?
Our best estimate based on the last decade of data is that the genetic component of intelligence is controlled by somewhere between 10,000 and 24,000 variants. We also know that each one, on average, contributes about +-0.2 IQ points.
Genetically altering IQ is more or less about flipping a sufficient number of IQ-decreasing variants to their IQ-increasing counterparts. This sounds overly simplified, but it’s surprisingly accurate; most of the variance in the genome is linear in nature, by which I mean the effect of a gene doesn’t usually depend on which other genes are present.
So modeling a continuous trait like intelligence is actually extremely straightforward: you simply add the effects of the IQ-increasing alleles to to those of the IQ-decreasing alleles and then normalize the score relative to some reference group.
To simulate the effects of editing on intelligence, we’ve built a model based on summary statistics from UK Biobank, the assumptions behind which you can find in the appendix
Based on the model, we can come to a surprising conclusion: there is enough genetic variance in the human population to create a genome with a predicted IQ of about 900. I don’t expect such an IQ to actually result from flipping all IQ-decreasing alleles to their IQ-increasing variants for the same reason I don’t expect to reach the moon by climbing a very tall ladder; at some point, the simple linear model will break down.
But we have strong evidence that such models function quite well within the current human range, and likely somewhat beyond it. So we should actually be able to genetically engineer people with greater cognitive abilities than anyone who’s ever lived, and do so without necessarily making any great trade-offs.
Even if a majority of iq-increasing genetic variants had some tradeoff such as increasing disease risk (which current evidence suggests they mostly don’t), we could always select the subset that doesn’t produce such effects. After all, we have 800 IQ points worth of variants to choose from!
Total maximum gain
Given the current set of publicly available intelligence-associated variants available in UK biobank, here’s a graph showing the expected effect on IQ from editing genes in adults.
Not very impressive! There are several factors at play deflating the benefits shown by this graph.
The limitations in 1-5 together reduce the estimated effect size by 79% compared to making perfect edits in an embryo. If you could make those same edits in an embryo, the gains would top out around 30 IQ points.
But the bigger limitation originates from the size of the data set used to train our predictor. The more data used to train an intelligence predictor, the more of those 20,000 IQ-affecting variants we can identify, and the more certain we can be about exactly which variant among a cluster is actually causing the effect.
And the more edits you can make, the better you can take advantage of that additional data. You can see this demonstrated pretty clearly in the graph below, where each line represents a differently sized training set.
Our current predictors are trained using about 135,000 samples, which would place it just above the lowest line on the graph. There are existing databases right now such as the million veterans project with sample sizes of (you guessed it) one million. The expected gain from editing using a predictor trained on such a data set is shown by the orange line in the graph above.
Companies like 23&Me genotyped their 12 millionth customer two years ago and could probably get at perhaps 3 million customers to take an IQ test or submit SAT scores. A predictor trained with that amount of data would perform about as well as the green line on the graph above.
So larger datasets could increase the effect of editing by as much as 13x! If we hold the number of edits performed constant at 2500, here’s how the expected gain would vary as a function of training set size:
Now we’re talking! If someone made a training set with 3 million data points (a realistic possibility for companies like 23&Me), the IQ gain could plausibly be over 100 points (though keep in mind the uncertainties listed in the numbered list above). Even if we can only make 10% of that many edits, the expected effect size would still be over one standard deviation.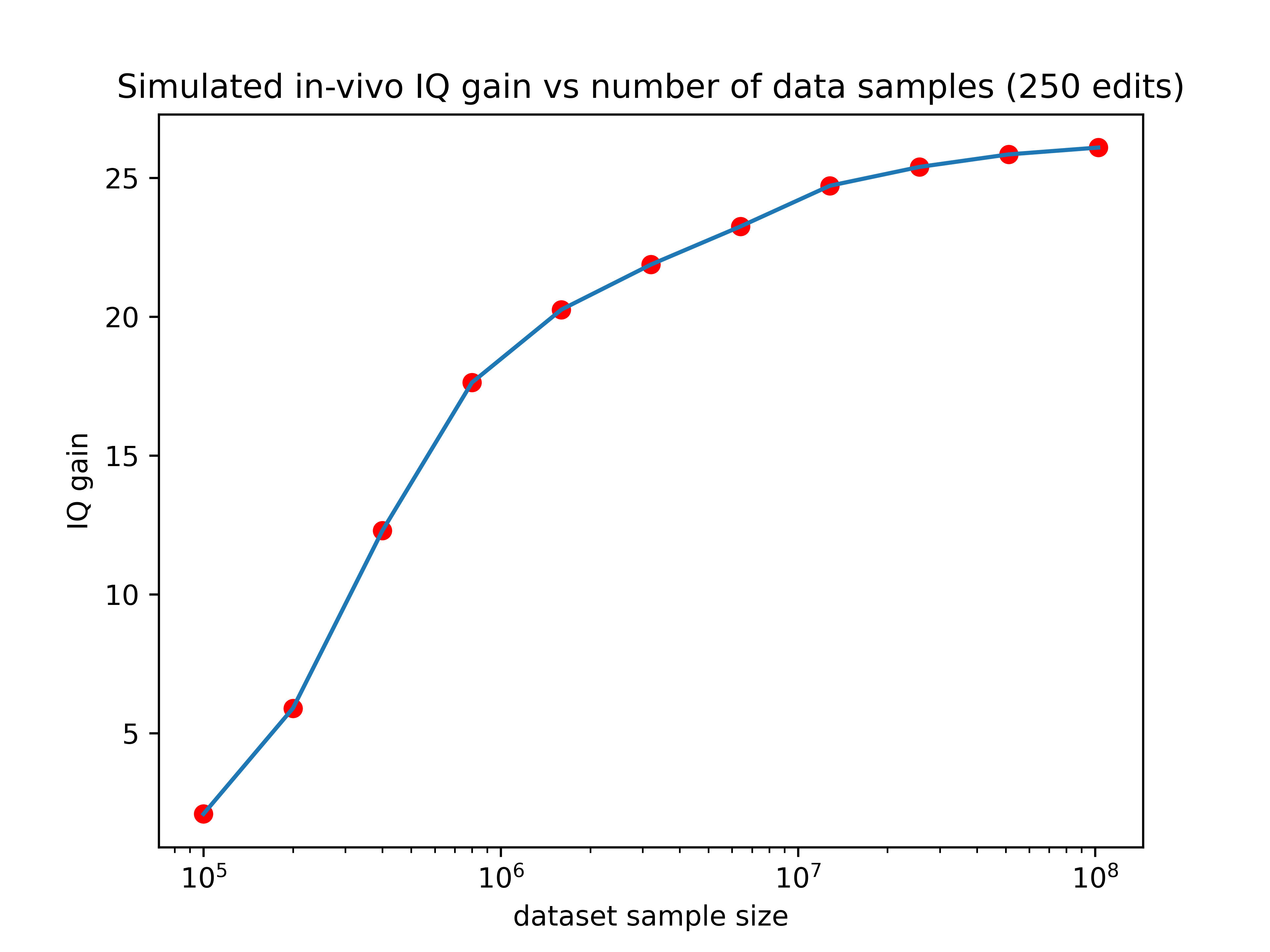
Lastly, just for fun, let’s show the expected effects of 2500 edits if they were made in an embryo and we didn’t have to worry about off-targets or limitations of editors (don’t take this graph too seriously)
The sample size issue could be solved pretty easily by any large government organization interested in collecting intelligence data on people they’ve already genotyped. However, most of them are not currently interested.
Why has it been so hard to get larger sample sizes?
There is a taboo against direct investigation of the genetics of intelligence in academia and government funding agencies. As a result, we have collected a pitifully small amount of data about intelligence, and our predictors are much worse as a result.
So people interested in intelligence instead have to research this proxy trait that is correlated with intelligence called “educational attainment”. Research into educational attainment IS allowed, so we have sample sizes of over 3 million genome samples with educational attainment labels and only about 130,000 for intelligence.
Government agencies have shown they could solve the data problem if they feel so inclined. As an alternative, if someone had a lot of money they could simply buy a direct-to-consumer genetic testing company and either directly ask participants to take an IQ test or ask them what their SAT scores were.
Or someone who wants to do this could just start their own biobank.
How can you even make these edits anyways? Isn’t CRISPR prone to errors?
Another of my initial doubts stemmed from my understanding of gene editing tools like CRISPR, and their tendency to edit the wrong target or even eliminate entire chromosomes.
You see, CRISPR was not designed to be a gene editing tool. It’s part of a bacterial immune system we discovered all the way back in the late 80s and only fully understood in 2012. The fact that it has become such a versatile gene editing tool is a combination of luck and good engineering.
The gene editing part of the CRISPR system is the Cas9 protein, which works like a pair of scissors. It chops DNA in half at a specific location determined by this neat little attachment called a “guide RNA”. Cas9 uses the guide RNA like a mugshot, bumping around the cell’s nucleus until it finds a particular three letter stretch of DNA called the PAM. Once it finds one, it unravels the DNA starting at the PAM and checks to see if the guide RNA forms a complementary match with the exposed DNA.
Cas9 can actually measure how well the guide RNA bonds to the DNA! If the hybridization is strong enough (meaning the base pairs on the guide RNA are complementary to those of the DNA), then the Cas9 “scissors” will cut the DNA a few base pairs to the right of the PAM.
So to replace a sequence of DNA, one must introduce two cuts: one at the start of the DNA and one at the end of it.
Once Cas9 has cut away the part one wishes removed, a repair template must be introduced into the cell. This template is used by the cell’s normal repair processes to fill in the gap.
Or at least, that’s how it’s supposed to work. In reality a significant portion of the time the DNA will be stitched back together wrong or not at all, resulting in death of the chromosome.
This is not good news for the cell.
So although CRISPR had an immediate impact on the life sciences after it was introduced in 2012, it was rightfully treated with skepticism in the context of human genome editing, where deleting a chromosome from someone’s genome could result in immediate cell death.
Had nothing changed, I would not be writing this post. But in 2016, a new type of CRISPR was created.
The first base editor
Many of the problems with CRISPR stem from the fact that it cuts BOTH strands of DNA when it makes an edit. A double stranded break is like a loud warning siren broadcast throughout the cell. It can result in a bunch of new base pairs being added or deleted at the break site. Sometimes it even results in the immune system ordering the cell to kill itself.
So when David Liu started thinking about how to modify CRISPR to make it a better gene editor, one of the first things he tried to do was prevent CRISPR from making double-stranded breaks.
He and members of his lab came up with a very creative way of doing this. They modified the Cas9 scissors so that they would still seek out and bind to a matching DNA sequence, but wouldn’t cut the DNA. They then attached a new enzyme to Cas9 called a “deaminase”. Its job is to change a single base through a chemical reaction.
They called this new version of CRISPR a “base editor”
There’s a lot more detail to go into regarding how base editors work which I will explain in another post, but the most noteworthy trait of base editors is how reliable they are in comparison to old school Cas9.
Base editors make the correct edit between 60 and 180 times as often as the original Cas9 CRISPR. They also make 92% fewer off-target edits and 99.7% fewer insertion and deletion errors.
The most effective base editor I’ve found so far is the transformer base editor, which has an editing efficiency of 90% in T cells and no detected off-targets.
But despite these considerable strength, base editors still have two significant weaknesses:
To address these issues, a few members of the Liu lab began working on a new project: prime editors.
Prime editors; the holy grail of gene editing technology?
Like base editors, prime editors only induce single stranded breaks in DNA, significantly reducing insertion and deletion errors relative to old school CRISPR systems. But unlike base editors, prime editors can change, insert, or delete up to about a hundred base pairs at a time.
The core insight leading to their creation was the discovery that the guide RNA could be extended to serve as a template for DNA modifications.
To greatly oversimplify, a prime editor cuts one of the DNA strands at the edit site and adds new bases to the end of it using an extension of the guide RNA as a template. This creates a bit of an awkward situation because now there are two loose flaps of DNA, neither of which are fully attached to the other strand.
Thankfully the repair enzyme in charge of fixing these types of problems (which are a necessary part of normal cellular function), really likes to cut the unedited flap for reasons I won’t get into. Once it has cut away the other flap and stitched together the edited strand, the situation looks like this:
There’s still one problem left to fix though: the unedited strand is… well… unedited.
To fix this, Liu and Co use another enzyme to cut the bottom strand of DNA near the edit site, which tricks the repair enzymes in charge of fixing mismatches into thinking that the unedited strand has been damaged and needs to be fixed so that it matches the edited one.
So the cell’s own repair enzymes actually complete the edit for us, which is pretty neat if you ask me.
Together, prime editors and base editors give us the power to edit around 90% of the genetic variants we’re interested in with very high precision. Without them, editing genes in a living human body would be infeasible due to off-target changes.
How do you even get editors into brain cells in the first place?
There are roughly 200 billion cells in an average brain. You can’t just use a needle to inject a bunch of gene editing tools into each one. So how do we get them inside?
There are several options available to us including viral proteins, but the delivery platform I like the most is called a lipid nanoparticle. Let me explain the idea.
Dual use technology
Remember COVID vaccines? The first ones released used this new technology called “mRNA”. They worked a little differently from old school immunizations.
The goal of a vaccine is to expose your immune system to part of a virus so that it will recognize and kill it in the future.
One way to do this is by taking a little piece of the virus and injecting it into your body. This is how the Novavax vaccine worked.
Another way is by sending an instruction to MAKE part of the virus to your body’s own cells. This is how the Pfizer and Moderna vaccines worked. And the instruction they used was a little piece of ribonucleic acid called mRNA.
mRNA is an instruction to your cells. If you can get it inside one, the cell’s ribosomes will transcribe mRNA into a protein. It’s a little bit like an executable file for biology.
By itself, mRNA is pretty delicate. The bloodstream is full of nuclease enzymes that will chop them into itty-bitty pieces.
Also, because mRNA basically has root level access to your cells, your body doesn’t just shuttle it around and deliver it like the postal service. That would be a major security hazard.
So to get mRNA inside a cell, COVID vaccines stick them inside a delivery container to protect them from the hostile environment outside cells and to get them where they need to go. This container is a little bubble of fat called a lipid nanoparticle.
If you inject these nanoparticles into your muscle tissue, they will float around for a while before being absorbed by your muscle cells via a process called endocytosis.
Afterwards, the mRNA is turned into a protein by the ribosomes in the muscle cell. In the case of COVID vaccines the protein is a piece of the virus called the “spike” that SARS-CoV-2 uses to enter your tissues.
Once created, some of these spike proteins will be chopped up by enzymes and displayed on the outside of each cell’s membrane. Immune cells patrolling the area will eventually bind to these bits of the spike protein and alert the rest of the immune system, triggering inflammation. This is how vaccines teach your body to fight COVID.
So here’s an idea: what if instead of making a spike protein, we deliver mRNA to make a gene editor?
Instead of manufacturing part of a virus, we’d manufacture a CRISPR-based editing tool like a prime editor. We could also ship the guide RNA in the same lipid nanoparticle, which would bond to the newly created editor proteins, allowing them to identify an editing target.
A nuclear localization sequence on the editor would allow the it to enter the cell’s nucleus where the edit could be performed.
So you inject lipid nanoparticles into the brain?
Not exactly. Brains are much more delicate than muscles, so we can’t just copy the administration method from COVID vaccines and inject nanoparticles directly into the brain with a needle.
Well actually we could try. And I plan to look into this during the mouse study phase of the project in case the negative effects are smaller than I imagine.
But apart from issues with possibly damaging brain or spinal tissue (which we could perhaps avoid), the biggest concern with directly injecting editors into the cerebrospinal fluid is distribution. We need to get the editor to all, or at least most cells in the brain or else the effect will be negligible.
If you just inject a needle into a brain, the edits are unlikely to be evenly distributed; most of the cells that get edited will be close to the injection site. So we probably need a better way to distribute lipid nanoparticles evenly throughout the brain.
Thankfully, the bloodstream has already solved this problem for us: it is very good at distributing resources like oxygen and nutrients evenly to all cells in the brain. So the plan is to inject a lipid-nanoparticle containing solution into the patient’s veins, which will flow through the bloodstream and eventually to the brain.
There are two additional roadblocks: getting the nanoparticles past the liver and blood brain barrier.
By default, if you inject lipid nanoparticles into the bloodstream, most of them will get stuck in the liver. This is nice if you want to target a genetic disease in that particular organ (and indeed, there have been studies that did exactly that, treating monogenic high cholesterol in macaques). But if you want to treat anything besides liver conditions then it’s a problem.
From my reading so far, it sounds like nanoparticles circulate through the whole bloodstream many times, and each time some percentage of them get stuck in the liver. So the main strategy scientists have deployed to avoid accumulation is to get them into the target organ or tissue as quickly as possible.
Here’s a paper in which the authors do exactly that for the brain. They attach a peptide called “angiopep-2” onto the outside of the nanoparticles, allowing them to bypass the blood brain barrier using the LRP1 receptor. This receptor is normally used by LDL cholesterol, which is too big to passively diffuse through the endothelial cells lining the brain. But the angiopep-2 molecule acts like a key, allowing the nanoparticles to use the same door.
The paper was specifically examining this for delivery of chemotherapy drugs to glial cells, so it’s possible we’d either need to attach additional targeting ligands to the surface of the nanoparticles to allow them to get inside neurons, or that we’d need to use another ligand besides angiopep. But this seems like a problem that is solvable with more research.
Or maybe you can just YOLO the nanoparticles directly into someone’s brain with a needle. Who knows? This is why research will need to start with cell culture experiments and (if things go well) animal testing.
What if administration via lipid nanoparticles doesn’t work?
If lipid nanoparticles don’t work, there are two alternative delivery vectors we can look into: adeno-associated viruses (AAVs) and engineered virus-like particles (eVLPs).
AAVs have become the go-to tool for gene therapy, mostly because viruses are just very good at getting into cells. The reason we don’t plan to use AAVs by default mostly comes down to issues with repeat dosing.
As one might expect, the immune system is not a big fan of viruses. So when you deliver DNA for a gene editor with an AAV, the viral proteins often trigger an adaptive immune response. This means that when you next try to deliver a payload with the same AAV, antibodies created during the first dose will bind to and destroy most of them.
There ARE potential ways around this, such as using a different version of an adenovirus with different epitopes which won’t be detected by antibodies. And there is ongoing work at the moment to optimize the viral proteins to provoke less of an immune response. But by my understanding this is still quite difficult and you may not have many chances to redose.
But there are other viral delivery vectors
Engineered virus-like particles, or eVLPs for short, are another way to get gene editors into cells. Like AAVs, they are constructed of various viral proteins and a payload.
But unlike AAVs, eVLPs usually contain proteins, not DNA. This is important because, once they enter a cell, proteins have a much shorter half-life than DNA. A piece of DNA can last for weeks or sometimes even months inside the nucleus. As long as it is around, enzymes will crank out new editor proteins using that DNA template.
Since most edits are made shortly after the first editors are synthesized, creating new ones for weeks is pointless. At best they will do nothing and at worst they will edit other genes that we don’t want changed. eVLPs fix this problem by delivering the editor proteins directly instead of instructions to make them.
They can also be customized to a surprising degree. The yellow-colored envelope glycoprotein shown in the diagram above can be changed to alter which cells take up the particles. So you can target them to the liver, or the lungs, or the brain, or any number of other tissues within the body.
Like AAVs, eVLPs have issues with immunogenicity. This makes redosing a potential concern, though it’s possible there are ways to ameliorate these risks.
Would edits to the adult brain even do anything?
Apart from the technical issues with making edits, the biggest uncertainty in this proposal is how large of an effect we should expect from modifying genes in adults. All the genetic studies on which our understanding of genes is based assume genes differ FROM BIRTH.
We know that some genes are only active in the womb, or in childhood, which should make us very skeptical that editing them would have an effect. For example, here’s a graph showing the relative level of expression of DCX, a gene active in neuronal progenitor cells and migrating immature neurons.
You can see the expression levels peak in the second trimester before dropping by a factor of ~80 by age 27.
We would therefore expect that any changes to promoters or inhibitors of the DCX gene would have little or no effect were they to be made in adulthood.
However, not every gene is like this. Here’s another graph from the same paper showing the relative expression of groups of genes in the hippocampus:
The red, blue and yellow lines, representing the expression of a group of synapse, dendrite, and myelination genes respectively, climb steadily until shortly after birth, then plateau throughout the rest of adulthood. So we could reasonably expect any changes to these genes to have a much stronger effect relative to changes to DEX.
This is one way to develop a basic prior about which genes are likely to have an effect if edited in adults: the size of the effect of a given gene at any given time is likely proportional to its level of expression. Genes that are only active during development probably won’t have as large of an effect when edited.
The paper I linked above collected information about the entire exome, which means we should be able to use its data to form a prior on the odds that editing any given variant will have an effect, though as usual there are some complications with this approach which I address in the appendix.
A systematic analysis of the levels of expressions of all genes throughout the lifespan is beyond the scope of this post, but beyond knowing some proteins reach peak levels in adulthood, here are a few more reasons why I still expect editing to have a large effect:
What if your editor targets the wrong thing?
Though base editors and prime editors have much lower rates of off-target editing than original CRISPR Cas9, off-target editing is still a concern.
There’s a particularly dangerous kind of off-target edit called a frameshift mutation that we really want to avoid. A frameshift mutation happens when you delete (or insert) a number of base pairs that is not a multiple of 3 from a protein-forming region.
Because base pairs are converted into amino acids three at a time, inserting or deleting a letter changes the meaning of all the base pairs that come after the mutation site.
This pretty much always results in a broken protein.
So to minimize the consequences of off-target mutations, we’d do better to target non-protein-coding regions. We already know quite well which regions of the genome are synthesized into proteins. And fortunately, it appears that 98% of intelligence-affecting variants are located in NON-CODING regions. So we should have plenty of variants to target.
What about off-target edits in non-coding regions?
Most of the insertions, deletions and disease-causing variants with small effects are in non-coding regions of DNA. This gives us some evidence that off-target edits made to these regions will not necessarily be catastrophic.
As long as we can avoid catastrophic off-target edits, a few off-targets will be ok so long as they are outweighed by edits with positive effects.
Part of the challenge here is characterizing off-target edits and how they are affected by choice of edit target. If your guide RNA’s binding domain ALMOST matches that of a really critical protein, you may want to think twice before making the edit. Whereas if it’s not particularly close to any target, you probably don’t need to worry as much.
However, it’s still unclear to me how big of a deal off-target edits and indels will be. If you unintentionally insert a few base pairs into the promoter region of some protein, will the promoter just work a little less well, or will it break altogether? I wasn’t able to find many answers in the research literature, so consider this a topic for future research.
It is worth pointing out that long-lived cells in the body such as neurons experience a thousand or more random mutations by age 40 and that, with the exception of cancer, these mutations rarely seem to cause serious dysfunction.
How about mosaicism?
Another potential issue with producing a large number of edits to cells in the brain is cellular mosaicism. The determination of which cell gets which set of edits is somewhat random. Maybe some cells are a little too far away from a blood vessel. Maybe some just don’t get enough of the editor. Regardless of the reason, it’s likely different cells within the brain will receive different sets of edits. Will this cause problems?
It’s possible it will. But as mentioned in the previous section, neurons in the adult brain already have significant genetic differences and most of people seem functional despite these variations. Neurons accumulate about 20-40 single letter mutations per year, with the exact amount varying by brain region and by individual.
By age 45 or so, the average neuron in an adult brain has about 1500 random genetic mutations. And the exact set of mutations each neuron has is somewhat random.
There are likely to be some differences between mutations caused by gene editing and those caused by cellular metabolism, pollutants, and cosmic rays. In particular, those caused by gene editing are more likely to occur at or near functionally important regions.
Still, it should be at least somewhat comforting that neurons usually remain functional when they have a few thousand genetic differences. But it is possible my optimism is misplaced here and mosaicism will turn out to be a significant challenge for some reason I can’t yet foresee. This is yet another reason to conduct animal testing.
Can we make enough edits in a single cell to have a significant effect?
Making a large enough number of edits to have a significant effect on a phenotype of interest is one of the main challenges with this research protocol. I’ve found fairly recent papers such as Qichen & Xue, that show a reasonably large number of edits can be made in the same cell at the same time; 31 edits using base editors and 3 using prime editors. As with nearly all research papers in this field, they used HEK293T cells, which are unusually easy to transfect with editors. We can therefore treat this as an upper bound on the number of simultaneous edits that can be made with current techniques.
Here’s another research paper from Ni et al. where the researchers used prime editors to make 8 simultaneous edits in regenerated wheat plants.
I also know of a PHD student in George Church’s lab that was able to make several thousand edits in the same cell at the same time by targeting a gene that has several thousand copies spread throughout the genome.
So people are trying multiplex editing, but if anyone has tried to make hundreds of edits at a time to different genes, they don’t seem to have published their results. This is what we need to be able to do for in-vivo multiplex editing to have a big impact on polygenic traits and diseases.
However, we don’t necessarily need to introduce all edits with a single injection. If the delivery vector and the editing process doesn’t provoke an adaptive immune response, or if immunosuppressants can be safely used many times, we can just do multiple rounds of administration; 100 edits during the first round, another 100 in the second round and so on.
If there is an immune response, there are some plausible techniques to avoid adverse reactions during subsequent doses, such as using alternative versions of CRISPR with a different antigenic profile. There are a limited number of these alternative CRISPRs, so this technique is not a general purpose solution.
Making hundreds of edits in cultured cells is the first thing I’d like to focus on. I’ve laid out my plans in more detail in “How do we make this real?”
Why hasn’t someone already done this?
There are hundreds of billions of dollars to be made if you can develop a treatment for brain disorders like Alzheimers, dementia and depression. There’s probably trillions to be made if you can enhance human intelligence.
And yet the people who have developed these tools seem to only be using them to treat rare monogenic disorders. Why?
Part of the answer is that “people are looking into it, but in a pretty limited capacity.” For example, here’s a paper that looked at multiplex editing of immune cells in the context of immunotherapy for cancer. And here’s a post that is strikingly similar to many of the ideas I’ve raised here. I only learned of its existence a week before publishing, but have spoken with the authors who had similar ideas about the challenges in the context of their focus on gene therapy for polygenic disease.
Apart from that I can’t find anything. There are 17 clinical trials involving some kind of CRISPR-based gene editing and I can’t find a single one that targets multiple genes.
So here I’m going to speculate on a couple of reasons why there seems to be so little interest in in-vivo multiplex editing.
Black box biology
To believe multiplex editing can work you have to believe in the power of black box biology. Unlike monogenic diseases, we don’t always understand the mechanism through which genetic variants involved in diabetes or intelligence cause their effects.
You have to believe that it is possible to affect a trait by changing genes whose effect you understand but whose mechanism of action you don’t. And what’s more, you need to believe you can change hundreds of genes in a large number of cells in this manner.
To most people, this approach sounds crazy, unpredictable, and dangerous. But the fact is nature mixes and matches combinations of disease affecting alleles all the time and it works out fine in most cases. There are additional challenges with editing such as off-targets and editing efficiency, but these are potentially surmountable issues.
Vague association with eugenics make some academics shy away
A lot of people in Academia are simply very uncomfortable with the idea that genes can influence outcomes in any context other than extremely simple areas like monogenic diseases. I recently attended a conference on polygenic embryo screening in Boston and the amount of denialism about the role of genetics in things as simple as disease outcomes was truly a sight to behold.
In one particularly memorable speech, Erik Turkheimer, a behavioral geneticist from University of Virginia claimed that, contrary to the findings of virtually every published study, adding more data to the training set for genetic predictors simply made them perform worse. As evidence to support this conclusion, he cited a university study on alcohol abuse with a sample size of 7000 that found no associations of genome-wide significance.
Turkheimer neglected to mention that no one could reasonably expect the study to unearth such associations because it was too underpowered to detect all but the most extreme effects. Most studies in this field have 30-100x the number of participants.
In many cases, arguing against these unsupported viewpoints is almost impossible. Genomic Prediction has published about a dozen papers on polygenic embryo selection, many of which are extremely well written and contain novel findings with significant clinical relevance. Virtually no one pays attention. Even scientists writing about the exact same topics fail to cite their work.
Many of them simply do not care what is true. They view themselves as warriors in a holy battle, and they already know anyone talking about genetic improvement is an evil racist eugenicist.
You can convince an open-minded person in a one-on-one discussion because the benefits are so gigantic and the goal is consistent with widely shared values as expressed in hundreds of other contexts.
But a large portion of Academics will just hear someone say “genes matter and could be improved”, think “isn’t that what Hitler did?”, and become forever unreachable. Even if you explain to them that no, Hitler’s theory of genetics was based on a corrupted version of purity ethics and that in fact a lot of his programs were probably dysgenic, and that even if they weren’t we would never do that because it’s evil, it’s too late. They have already passed behind the veil of ignorance. The fight was lost the moment the association with Hitler entered their mind.
Though these attitudes don’t completely prevent work on gene therapies, I think it likely that the hostility towards altering outcomes through genetic means drives otherwise qualified researchers away from this line of work.
Maybe there’s some kind of technical roadblock I’m missing
I’ve now talked to some pretty well-qualified bio PHDs with expertise in stem cells, gene therapy, and genetics. While many of them were skeptical, none of them could point to any part of the proposed treatment process that definitely won’t work. The two areas they were most skeptical of was the ability to deliver an editing vector to the brain and the ability to perform many edits in the same cell. But both seem addressable. Delivery for one, may be solved by other scientists trying to deliver gene editors to the brain to target monogenic conditions like Huntington’s.
There are unknowns of course, and many challenges which could individually or in conjunction make the project infeasible. For example either of the challenges listed above, or perhaps mosaicism is a bigger deal than I expect it to be.
But I’ve only talked to a couple of people. Maybe someone I haven’t spoken with knows of a hard blocker I am unaware of.
People are busy picking lower hanging fruit
It’s a lot easier to target a specific gene that causes a particular disease than it is to target hundreds or thousands of genes which contribute to disease risk. Given prime editors were only created four years ago and base editors only seven, maybe the people with the relevant expertise are just busy picking lower hanging fruit which has a much better chance of resulting in a money-making therapy. The lowest hanging fruit is curing monogenic diseases, and that’s currently where all the focus is.
The market is erring
Some types of markets are highly efficient. Commodities and stocks, for example, are very efficient because a large number of people have access to the most relevant information and arbitrage can happen extremely quickly.
But the more time I’ve spent in private business, and startups in particular, the more I’ve realized that most industries are not as efficient as the stock market. They are far less open, the knowledge is more concentrated, the entry costs are higher, and there just aren’t that many people with the abilities needed to make the markets more efficient.
In the case of multiplex editing, there are many factors at play which one would reasonably expect to result in a less efficient market:
How many people in the world satisfy all five of those criteria? My guess is not many. So despite how absurd it sounds to say that the pharmaceutical industry is overlooking a potential trillion dollar opportunity, I think there is a real possibility they are. Base editors have been around for seven years and in that time the number of papers published on multiplex editing in human cells for anything besides cancer treatment has remained in the single digits.
I don’t think this is a case of “just wait for the market to do it”.
Can we accelerate development of this technology?
After all that exposition, let me summarize where the work I think will be done by the market and that which I expect them to ignore.
Problems I expect the market to work on
Problems the market might solve
If you are targeting a monogenic disease, you only need to deliver an editing vector once. The adaptive immune response to the delivery vector and CRISPR proteins is therefore of limited concern; the innate immune response to these is much more important.
There may be overlap between avoiding each of these, but I expect that if an adaptive immune response is triggered by monogenic gene therapy and it doesn’t cause anaphylaxis, companies will not concern themselves with avoiding it.
The odds of making an off-target edit scale exponentially as the number of edits increase. Rare off-targets are therefore a much smaller concern for monogenic gene therapy, and not much of a concern at all for immunotherapy unless they cause cancer or are common enough to kill off a problematic number of cells.
I therefore don’t expect companies or research labs to address this problem at the scale it needs to be addressed within the multiplex editing context.
One possible exception are companies using gene editing in the agricultural space, but the life of a wheat plant or other crop has virtually zero value, so the consequences of off-targets are far less serious.
Problems I don’t expect the market to work on
Unless embryo selection for intelligence takes off, I think it is unlikely any of the groups that currently have the data to create strong intelligence predictors (or could easily obtain it) will do so.
If you don’t intend to target polygenic disease, there’s little motivation to make multiple genetic changes in cells. And as explained above, I don’t expect many if any people to work on polygenic disease risk because mainstream scientists are very skeptical of embryo selection and of the black box model of biology.
None of the monogenic diseases targeted by current therapies involve genes that are only active during childhood. It therefore seems fairly unlikely that anyone will do research into this topic since it has no clear clinical relevance.
How do we make this real?
The hardest and most important question in this post is how we make this technology real. It’s one thing to write a LessWrong post pointing out some revolutionary new technology might be possible. It’s quite another to run human trials for a potentially quite risky medical intervention that doesn’t even target a recognized disease.
Fortunately, some basic work is already being done. The first clinical trials of prime editors are likely to start in 2024. And there are at least four clinical trials underway right now testing the efficacy of base editors designed to target monogenic diseases. The VERVE-101 trial is particularly interesting due to its use of mRNA delivered via lipid nanoparticles to target a genetic mutation that causes high cholesterol and cardiovascular disease.
If this trial succeeds, it will definitively demonstrate that in-vivo gene editing with lipid nanoparticles and base editors can work. The remaining obstacles at that point will be editing efficiency and getting the nanoparticles to the brain (a solution to which I’ve proposed in the section titled “So you inject lipid nanoparticles into the brain?”. Here’s an outline of what I think needs to be done, in order:
The goal of such trials would be to test our hypotheses about mosaicism, cancer, and the relative effect of genes in adulthood vs childhood.
It’s possible one or more steps could be skipped, but at the very least you need to run trials in some kind of model organism to refine the protocol and ensure the treatment is safe enough to begin human trials.
My current plan
I don’t have a formal background in biology. And though I learn fairly quickly and have great resources like SciHub and GPT4, I am not going to be capable of running a lab on my own any time soon.
So at the moment I am looking for a cofounder who knows how to run a basic research lab. It is obviously helpful if you have worked specifically with base or prime editors before but what I care most about is finding someone smart that I get along with who will persevere through the inevitable pains of creating a startup. If anyone reading this is interested in joining me and has such a background, send me an email.
I’m hoping to obtain funding for one year worth of research, during which our main goal will be to show we can perform a large number of distinct edits in cell culture. If we are successful in a year the odds of us being able to modify polygenic brain based traits will be dramatically increased.
At that point we would look to either raise capital from investors (if there is sufficient interest), or seek a larger amount of grant funding. I’ve outlined a brief version of additional steps as well as a rough estimate of costs below.
Short-term logistics concerns
A rough outline of my research agenda
Rough estimate of costs
2000
Conclusion
Someone needs to work on massive multiplex in-vivo gene editing. This is one of the few game-board-flipping technologies that could have a significant positive impact on alignment prior to the arrival of AGI. If we completely stop AI development following some pre-AGI disaster, genetic engineering by itself could result in a future of incredible prosperity with far less suffering than exists in the world today.
I also think it is very unlikely that this will just “happen anyways” without any of us making an active effort. Base editors have been around since 2016. In that time, anyone could have made a serious proposal and started work on multiplex editing. Instead we have like two academic papers and one blog post.
In the grand scheme of things, it would not take that much money to start preliminary research. We could probably fund myself and my coauthor, a lab director, and lab supplies to do research for a year for under a million dollars. If you’re interested in funding this work and want to see a more detailed breakdown of costs than what I’ve provided above, please send me an email.
I don’t know exactly how long it will take to get a working therapy, but I’d give us a 10% chance of having something ready for human testing within five years. Maybe sooner if we get more money and things go well.
Do I think this is more important than alignment work?
No.
I continue to think the most likely way to solve alignment is for a bunch of smart people to work directly on the problem. And who knows, maybe Quintin Pope and Nora Belrose are correct and we live in an alignment by default universe where the main issue is making sure power doesn’t become too centralized in a post-AGI world.
But I think it’s going to be a lot more likely we make it to that positive future if we have smarter, more focused people around. If you can help me make this happen through your expertise or funding, please send me a DM or email me.
Appendix (Warning: boredom ahead)
Additional headaches not discussed
Regarding editing
Bystander edits
If you have a sequence like “AAAAA” and you’re trying to change it to “AAGAA”, a base editor will pretty much always edit it to “GGGGG”. This is called “bystander editing” and will be a source of future headaches. It happens because the deaminase enzyme is not precise enough to only target exactly the base pair you are trying to target; there’s a window of bases it will target. When I’ve written some software to identify which genetic variants we want to target I will be able to give a better estimate of how many edits we can target with base editors without inducing bystander edits. There are papers that have reduced the frequency of bystander edits, such as Gehrke et al, but it’s unclear to me at the moment whether the improvement is sufficient to make base editors usable on such sequences.
Chemical differences of base editor products
Traditional base editors change cytosine to uracil and adenine to inosine. Many enzymes and processes within the cell treat uracil the same as thymine and inosine the same as guanine. This is good! It’s why base editors work in the first place. But the substituted base pairs are not chemically identical.
It’s not clear to me at this moment what the implications of this substitution are. Given there are four clinical trials underway right now which use base editors as gene therapies, I don’t think they’re a showstopper; at most, I think they would simply reduce editing efficiency. But these chemical differences are something we will need to explore more during the research process.
Editing silenced DNA
There is mixed information online about the ability of base and prime editors to change DNA that is methylated or wrapped around histones. Some articles show that you can do this with modified base editors, but imply normal base editors have limited ability to edit such DNA. Also the ability seems to depend on the type of chromatin and the specific editor.
Avoiding off-target edits
Base editors (and to a lesser extent prime editors) have issues with off-target edits. The rate of off-targets depends on a variety of factors, but one of the most important is the similarity between the sequence of the binding site and that of other sites in the genome. If the binding site is 20 bases long, but there are several other sequences in the genome that only differ by one letter, there’s a decent chance the editor will bind to one of those other sequences instead.
But quantifying the relationship between the binding sequence and the off-target edit rate is something we still need to do for this project. We can start with models such as Deepoff, DeepCRISPR, Azimuth/Elevation, and DeepPrime to select editing targets (credit to this post for bringing these to my attention), then validate the results and possibly make adjustments after performing experiments.
Regarding fine-mapping of causal variants
There’s an annoying issue with current data on intelligence-affecting variants that reduces the odds of us editing the correct variant. As of this writing, virtually all of the studies examining the genetic roots of intelligence used SNP arrays as their data source. SNP arrays are a way of simplifying the genome sequencing process by focusing on genes that commonly vary between people. For example, the SNP array used by UK Biobank measures all genetic regions where at least ~1% of the population differs.
This means the variants that are implicated as causing an observed intelligence difference will always come from that set, even if the true causal variant is not included in the SNP array. This is annoying, because it sometimes means we’ll misidentify which gene is causing an observed difference in intelligence.
Fortunately there’s a relatively straightforward solution: in the short run we can use some of the recently released whole genome sequences (such as the 500k that were recently published by UK biobank) to adjust our estimates of any given variant causing the observed change. In the long run, the best solution is to just retrain predictors using whole genome data.
Regarding differential gene expression throughout the lifespan
Thanks to Kang et al. we have data on the level of expression of all proteins in the brain throughout the lifespan. But for reasons stated in “What if your editor targets the wrong thing”, we’re planning to make pretty much all edits to non-protein coding regions. We haven’t yet developed a mechanistic plan for linking non-coding regions with the proteins whose expression they affect. There are some cases where this should be easy, such as tying a promoter (which is always directly upstream of the exonic region) to a protein. But in other cases such as repressor genes which aren’t necessarily close to the affected gene, this may be more difficult.
Regarding immune response
I am not currently aware of any way to ensure that ONLY brain cells receive lipid nanoparticles. It’s likely they will get taken up by other cells in the body, and some of those cells will express editing proteins and receive edits. These edits probably won’t do much if we are targeting genes involved in brain-based traits, but non-brain cells do this thing called “antigen presentation” where they chop up proteins inside of themselves and present them on their surfaces.
It’s plausible this could trigger an adaptive immune response leading to the body creating antibodies to CRIPSR proteins. This wouldn’t necessarily be bad for the brain, but it could mean that subsequent dosings would lead to inflammation in non-brain regions. We could potentially treat this with an immunosupressant, but then the main advantage of lipid nanoparticles being immunologically inert kind of goes away.
Regarding brain size
Some of the genes that increase intelligence do so by enlarging the brain. Brain growth stops in adulthood, so it seems likely that any changes to genes involved in determining brain size would simply have no effect in adults. But it has been surprisingly hard to find a definitive answer to this question.
If editing genes involved in brain growth actually restarted brain growth in adulthood AFTER the fusion of growth plates in the cranium, it would cause problems.
If such a problem did occur we could potentially address it by Identifying a set of genes associated with intelligence and not brain size and exclusively edit those. We have GWAS for brain size such as this one, though we would need to significantly increase the data set size to do this.
But this is a topic we plan to study in more depth as part of our research agenda because it is a potential concern.
Editing induced amnesia?
Gene editing in the brain will change the levels and timings of various proteins. This in turn will change the way the brain learns and processes information. This is highly speculative, but maybe this somehow disrupts old memories in a way similar to amnesia.
I’m not very concerned about this because the brain is an incredibly adaptable tissue and we have hundreds of examples of people recovering functionality after literally losing large parts of their brain due to stroke or trauma. Any therapeutic editing process that doesn’t induce cancer or break critical proteins is going to be far, far less disruptive than traumatic brain injuries. But it’s possible there might be a postoperative recovery period as people’s brains adjust to their newfound abilities.
An easy way to de-risk this (in addition to animal trials) is by just doing a small number of edits during preliminary trials of intelligence enhancement.
Model assumptions
The model for gain as a function of number of edits was created by Kman with a little bit of feedback from GeneSmith.
It makes a few assumptions:
This is obviously not a very realistic model, but it probably produces fairly realistic results. But again, this is an area for future improvement.
Editor properties
Here’s a table Kman and I have compiled showing a good fraction of all the base and prime editors we’ve discovered with various attributes.
N/A
If you’ve made it this far send me a DM. I want to know what motivates people to go through all of this.